First Login & Environment Setup
First login
If you are participating in this course with a teacher, you have received a link and a password. Copy-paste the link (including the port, e.g.: http://12.345.678.91:10002) in your browser. This should result in the following page:
The link gives you access to a web version of Visual Studio Code. This is a powerful code editor that you can also use as a local application on your computer.
Type in the password that was provided to you by the teacher. Now let’s open the terminal. You can do that by clicking Application menu > Terminal > New Terminal:
For a.o. efficiency and reproducibility it makes sense to execute your commands from a script. With use of the ‘new file’ button:
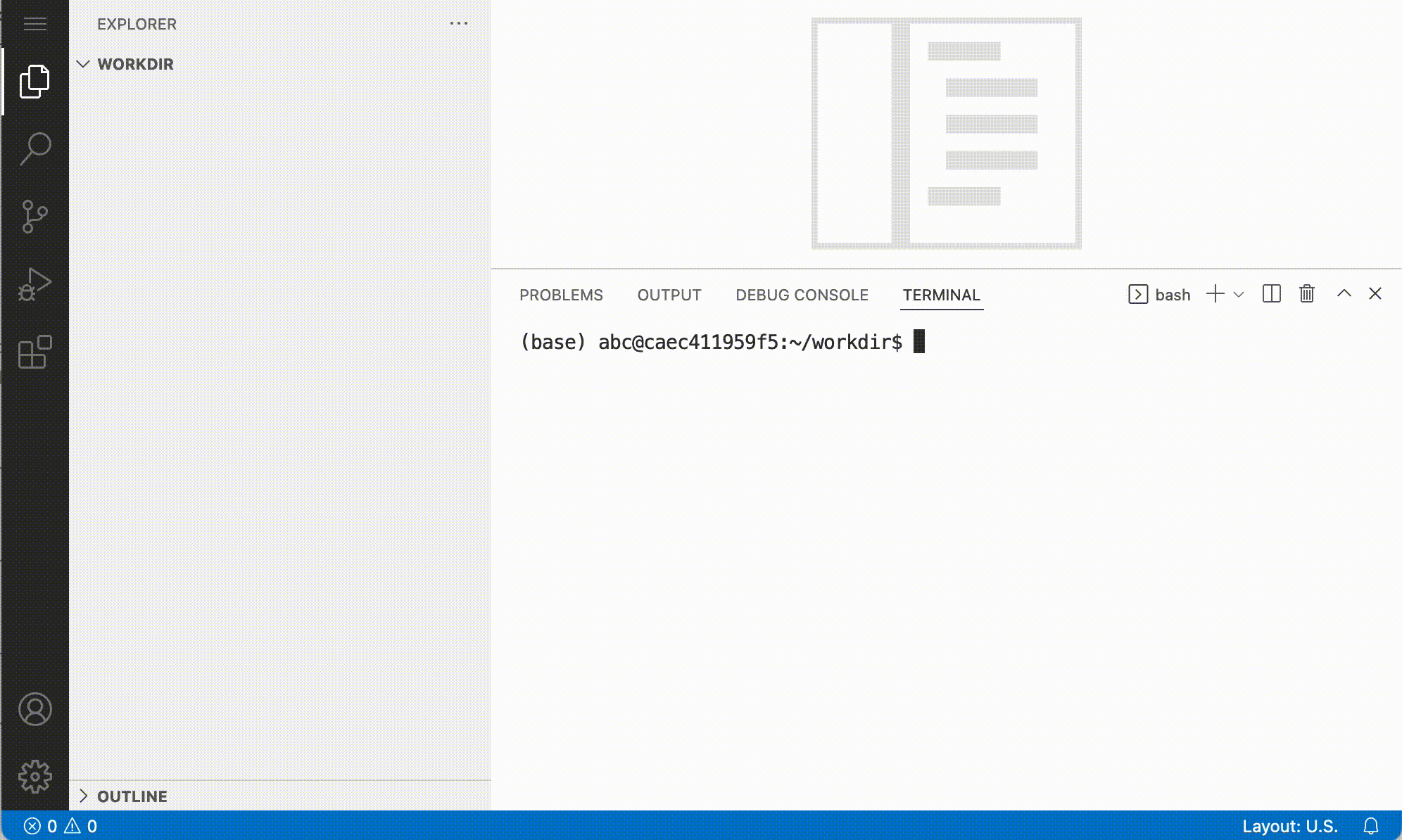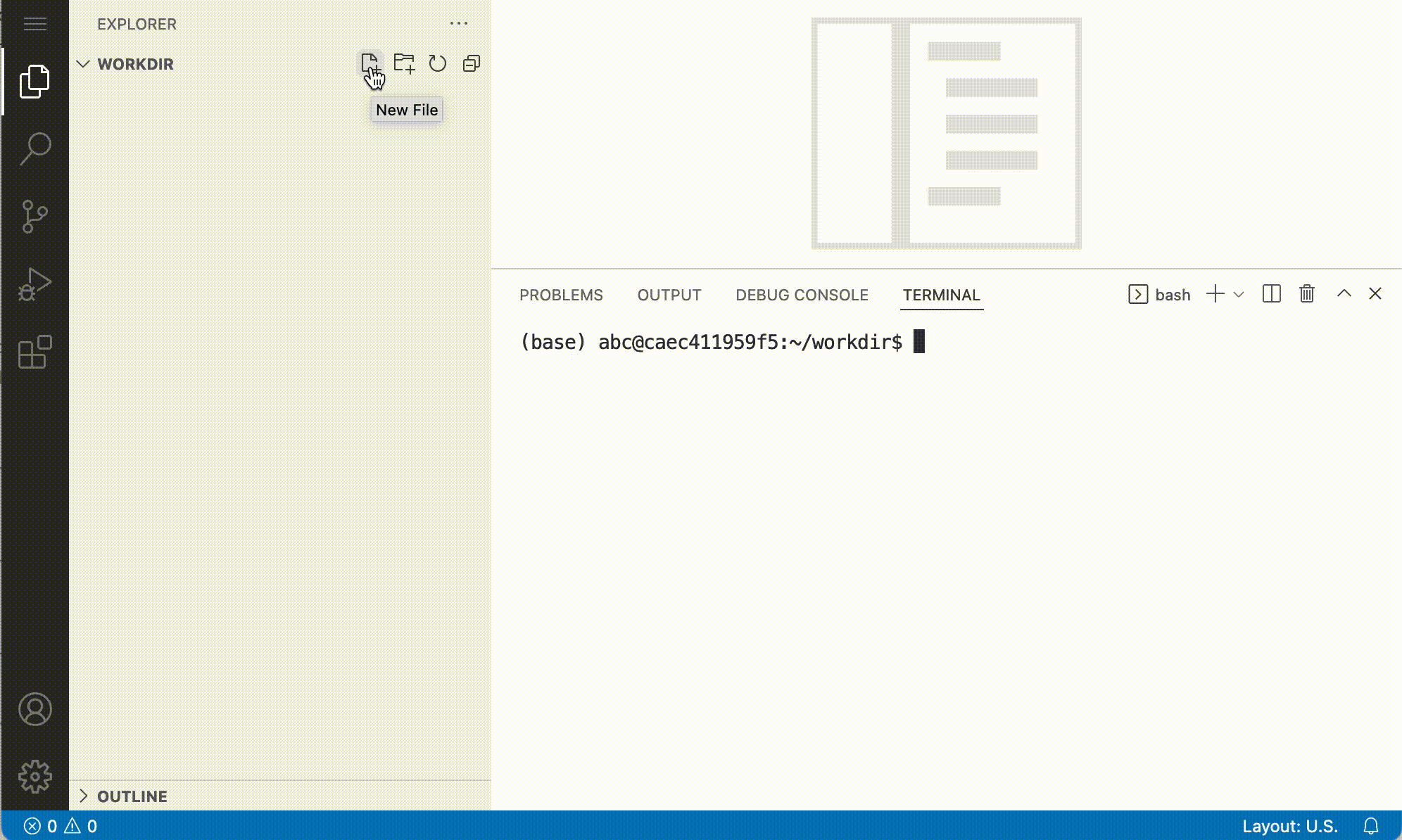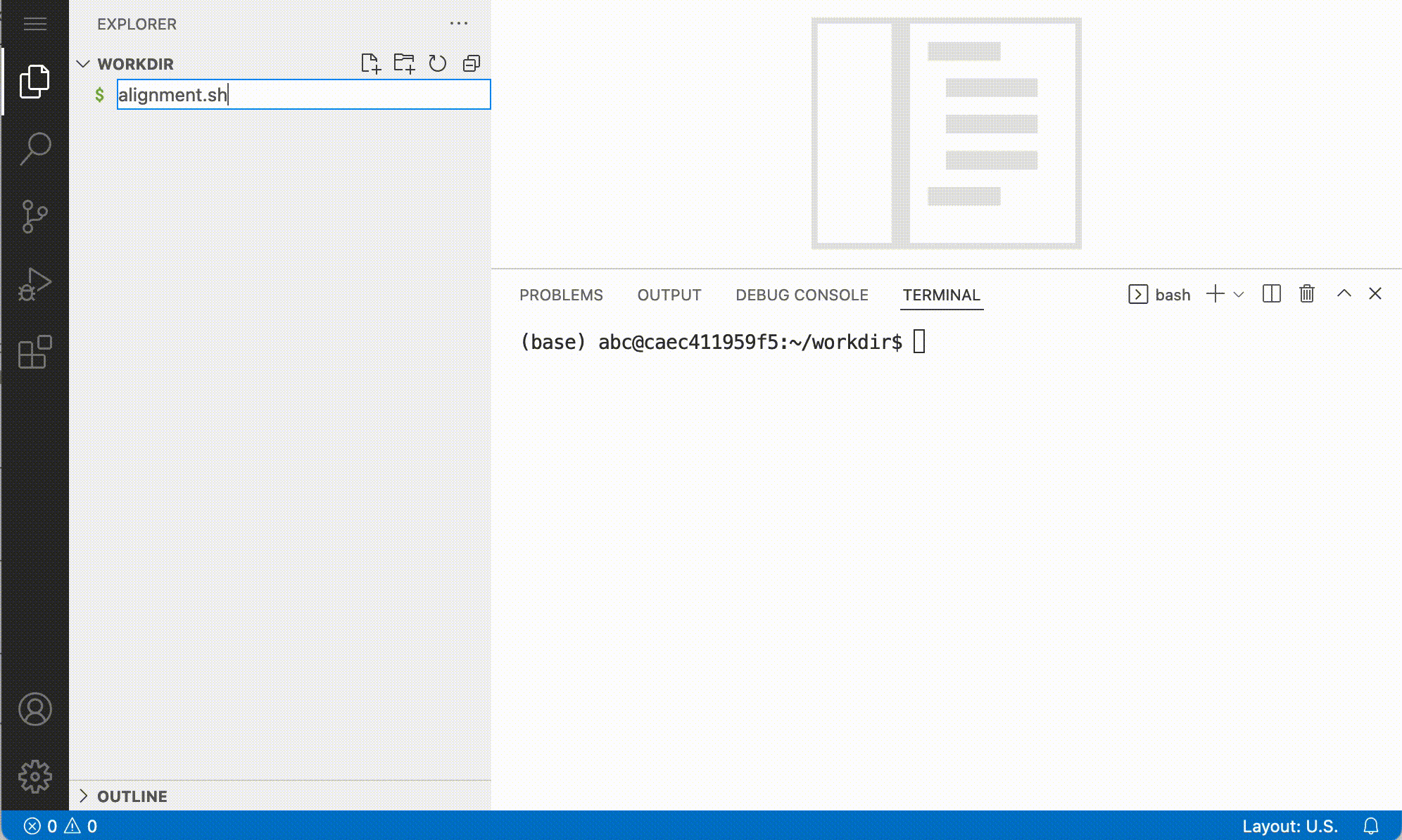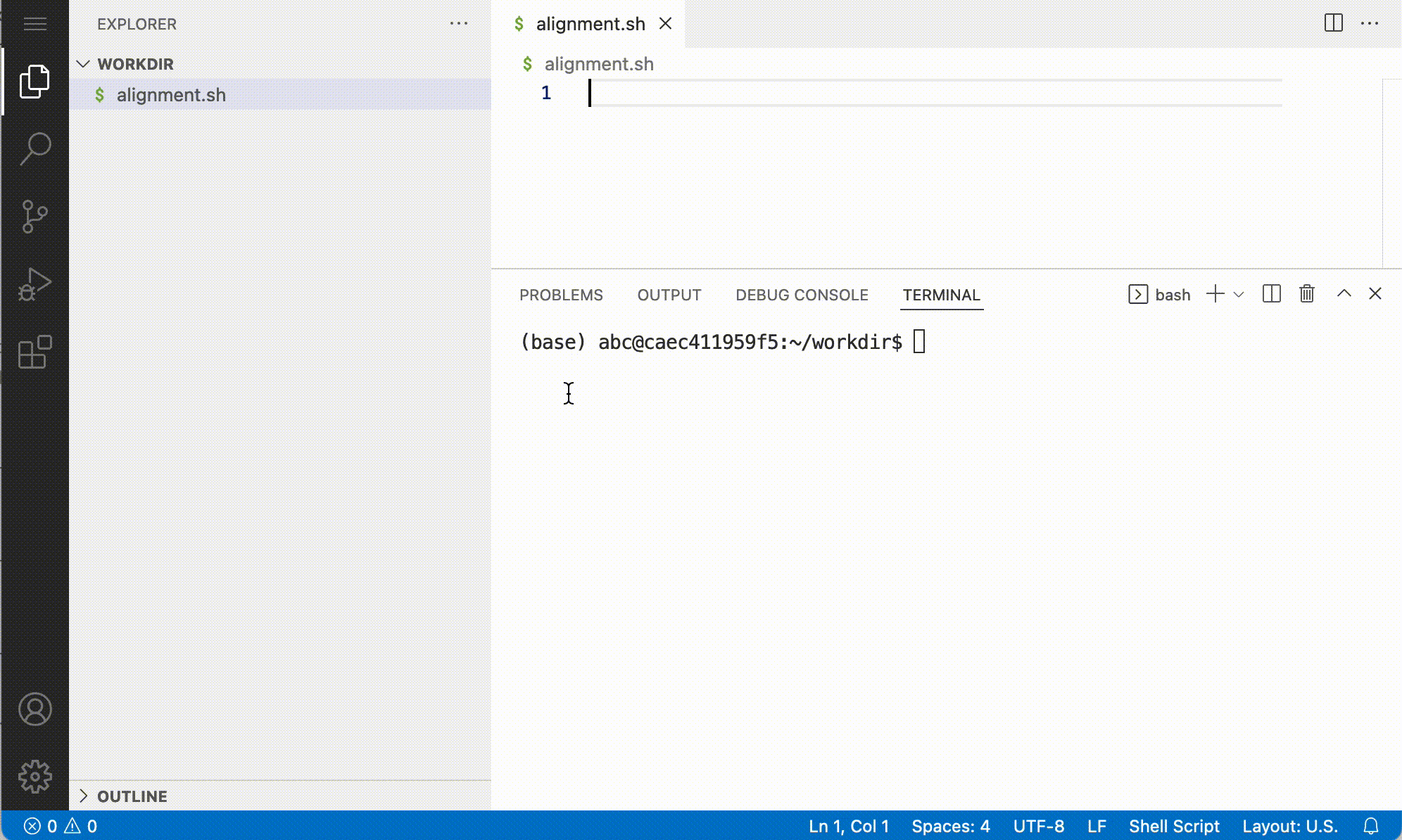
Setup
We will start the exercises with pre-aligned bam files. We have already prepared the folders for you with all the data that you need. If you are following the course later on:
You can create a symbolic link to the parent /data directory into your project directory project/, this simplify the path organization
cd ~/project
rmdir data
ln -s /data .This allows you to browse the data using VSCode’s file explorer within your project.
To start, download and extract to the into a folder, and change your paths to your folder:
mkdir -p ~/project
cd ~/project
wget https://cancer-variants-training.s3.eu-central-1.amazonaws.com/course_data.tar.gz
tar -xvzf course_data.tar.gz
mv course_data data
rm course_data.tar.gzThen update your paths, depending on where your data is:
PROJECT=~/project
ALIGNDIR=~/project/data/alignments
REFDIR=~/project/data/reference
RESOURCEDIR=~/project/data/resources
VEPDBS=~/project/data/VEP_dbsNow, check out the directory ~/project/data and see what’s in there (e.g. with tree):
└── ~/project/data
├── alignments
│ ├── normal.recal.bai
│ ├── normal.recal.bam
│ ├── tumor.recal.bai
│ └── tumor.recal.bam
├── reads
│ ├── normal_R1.fastq.gz
│ ├── normal_R2.fastq.gz
│ ├── tumor_R1.fastq.gz
│ └── tumor_R2.fastq.gz
├── reference
│ ├── exome_regions.bed.interval_list
│ ├── ref_genome.dict
│ ├── ref_genome.fa
│ ├── ref_genome.fa.fai
│ └── ref_genome.fa.gz
├── resources
│ ├── 1000G_phase1.snps.high_confidence.hg38.subset.vcf.gz
│ ├── 1000G_phase1.snps.high_confidence.hg38.subset.vcf.gz.tbi
│ ├── 1000G_phase1.snps.high_confidence.hg38.vcf.gz
│ ├── 1000G_phase1.snps.high_confidence.hg38.vcf.gz.tbi
│ ├── 1000g_pon.hg38.subset.vcf.gz
│ ├── 1000g_pon.hg38.subset.vcf.gz.tbi
│ ├── 1000g_pon.hg38.vcf.gz
│ ├── 1000g_pon.hg38.vcf.gz.tbi
│ ├── af-only-gnomad.hg38.subset.vcf.gz
│ ├── af-only-gnomad.hg38.subset.vcf.gz.tbi
│ ├── af-only-gnomad.hg38.vcf.gz
│ ├── af-only-gnomad.hg38.vcf.gz.tbi
│ ├── Mills_and_1000G_gold_standard.indels.hg38.subset.vcf.gz
│ ├── Mills_and_1000G_gold_standard.indels.hg38.subset.vcf.gz.tbi
│ ├── Mills_and_1000G_gold_standard.indels.hg38.vcf.gz
│ ├── Mills_and_1000G_gold_standard.indels.hg38.vcf.gz.tbi
│ ├── refFlat.txt
│ └── refFlat.txt.gz.1
└── VEP_dbs
├── alphamissense
│ ├── AlphaMissense_hg38.tsv.gz
│ └── AlphaMissense_hg38.tsv.gz.tbi
├── clinvar
│ ├── clinvar.vcf.gz
│ └── clinvar.vcf.gz.tbi
└── revel
├── new_tabbed_revel_grch38.tsv.gz
├── new_tabbed_revel_grch38.tsv.gz.tbi
└── new_tabbed_revel.tsv.gz
9 directories, 38 filesShowing us that we have four directories:
alignments: containing bam files of tumor and normalreference: containing the genome fasta file and target intervalsresources: containing amongst other variant files (vcf) from among other the 1000 genomes project and gnomADVEP_dbs: Clinvar and alphamissense and revel databases for usage with VEP.
The dataset we’re working with is prepared by the developers of the Precision medicine bioinformatics course by the Griffith lab. It is whole exome sequencing (WES) data of cell lines derived from a tumor of triple negative breast cancer (TNBC) (HCC1395) and derived from normal tissue (HCC1395 BL).
You are strongly encouraged to your work with scripts during the course, which you store in the directory scripts. Therefore create a scripts directory:
cd ~/project/
mkdir scripts